| Calculate Intersections |
The general idea general
During this stage intersections between links are calculated and added as new nodes into the graph structure. In the end this will lead to a graph describing enclosed areas that do not overlap each other, in order to determine for each area the boolean end result.
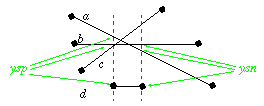
The intersections are found using scanbeam algorithms ( See Scanbeams. ). Snapping nodes to lines and nodes to nodes is an integrated part of this. The whole procedure to calculate intersections looks to much complicated. But this is all due to the integrated snapping feature, which is essential to reach a stable algorithm that can be used for extremely complicated drawings. Snapping a node to a link is done by looking at the distance in vertical and horizontal direction, because this can be done quickly. The effect is that in reality we check if the node is within 1/sqrt(MARGE) distance to the link, this is acceptable. intersecting links steps. gives an example of the steps involved

The end result is a bigger graph, since links are added to it. All of this is happening during the process of scanning the graph with a scanbeam. At the moment a link is removed from the scanbeam (at the most to the left node of this link, which is the _low of the current scanbeam), the link is checked for interscetions.
If there are interscetion with this link, they are inserted into the graph directly, the new links generated because of this are inserted at the beginning of the graph (sorted list of links), so they will not effect the rest of the graph that is more to the right and still needs to be scanned.
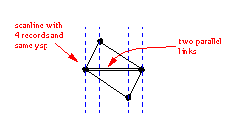
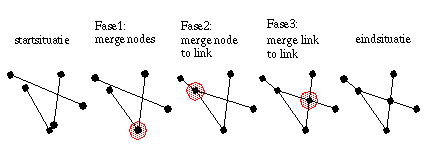
When nodes are within MARGE distance to eachother, they are merged into one node. For this the links are sorted on the X and Y of the beginnode. The bin flag in the link is used to tell if that beginnode is already processed. Using the sort information we can merge the beginnodes of coming links to the beginnode of the current link. Merging nodes can result in links that are reduced to zero lenght, they are removed at the end.
Since nodes are moved in position during a merge of two nodes, there is always a change that new intersections between links will occur. This is the case in general, snapping nodes to lines also may cause extra intersection somewhere else in the graph. This is why the snapping process is not integrated with the calculation of the intersections between links (Link to Link), and also that (Node to Link) is after (Node to Node). This sequence of steps minimizes the change of generating weird intersections.
The whole idea is to generate a graph inclusif all intersections, in order to form enclosed areas that do not and may not overlap. If because of bad calculation of intersections, those enclosed areas do overlap, the boolean result will be wrong or even fail.
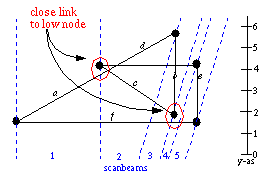
For each scanbeam all the records are checked, to see if the link of the record
is within MARGE distance to the low node (only in vertical direction). Only
the low node needs to be checked, since this is the only new node inserted in
the new beam. If the link of a record has as the begin or end node the low itself
it will be skipped. If a node is too close to a link, the node is put in the
crosslist of the link (the crosslist is maintained as part of the each link).
The nodes put into the crosslist will be processes if the link is removed from
the beam, it will split the link into several links containing the nodes.
Checking for close links to the current low node is done at the moment new links
are inserted in the beam or when old links are removed from the beam. Every
node in the graph will be checked this way, since every beginnode of a link
is the endnode of another link. This is because the input is a set of polygons,
so looping.

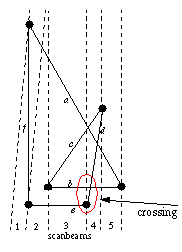
The following cases can be optimized: